Extending Dynamic Flux Balance Modeling to Feedback-Regulated Modular Co-Culture Systems
- Rutuja Bhamare
- Mar 2
- 2 min read
There exists an experimentally motivated co-culture framework validated in situ by Timothy J. Hanly, Morgan Urello, and Michael A. Henson in their study published in Applied Microbiology and Biotechnology on “Dynamic flux balance modeling of S. cerevisiae and E. coli co-cultures for efficient consumption of glucose/xylose mixtures,” which integrates dynamic flux balance analysis (dFBA) with bioreactor modeling to describe substrate utilization in S. cerevisiae and E. coli co-cultures.
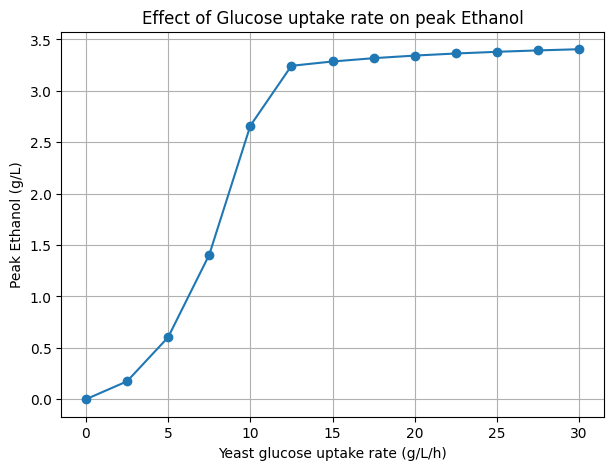
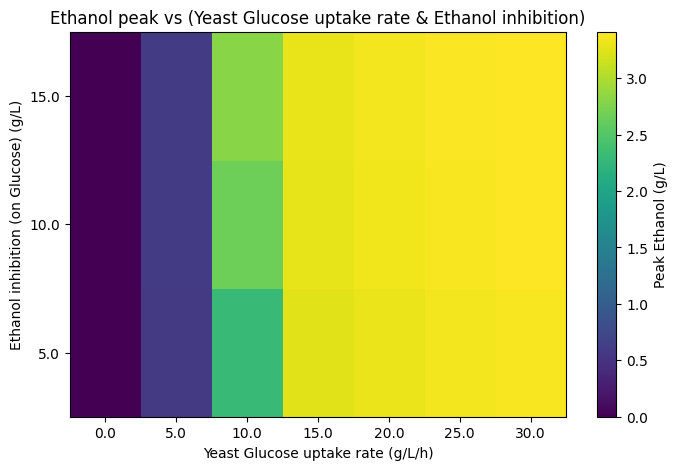
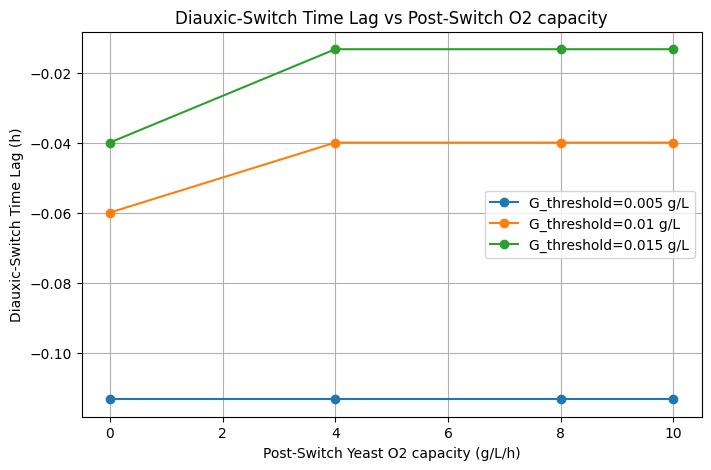
In replicating this work, we reconstructed the dFBA framework to simulate glucose and xylose consumption dynamics and ethanol production under varying environmental and kinetic constraints. Our reproduced simulations successfully captured the characteristic diauxic behavior observed in the original study, with yeast preferentially consuming glucose before shifts in metabolic activity altered overall substrate partitioning. Through systematic parameter exploration, we examined the sensitivity of ethanol accumulation and identified that elevated ethanol concentrations arise from high glucose uptake rates combined with oxygen limitation and insufficient dilution or downstream metabolic consumption. Furthermore, diauxic timing was found to be strongly controlled by substrate depletion thresholds and oxygen transfer capacity, confirming the system’s sensitivity to environmental and transport parameters.
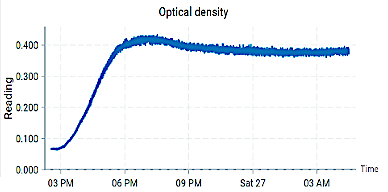
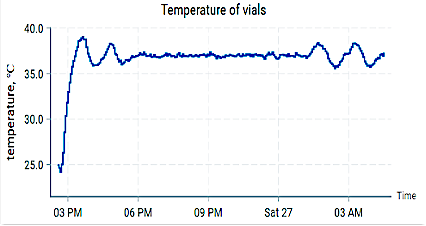
These findings provide a mechanistic foundation for the modular bioreactor platform currently being developed in our laboratory, where yeast and E. coli are maintained in physically separated but metabolically coupled reactors. Unlike traditional mixed co-cultures, this configuration enables controlled metabolite exchange while preserving independent environmental regulation. Our reinforcement-learning–based control architecture, implemented on a simplified metabolic network, dynamically adjusts feed and circulation rates to sustain desired growth conditions. Within the experimental setup, optical density (OD) and temperature are continuously monitored to ensure reliable growth quantification and stable kinetic conditions across vials. OD measurements serve as real-time proxies for biomass concentration, enabling direct comparison between predicted and observed growth dynamics, while temperature regulation and tracking support reproducibility and consistent metabolic performance. By integrating adaptive control with model-informed design, this modular setup bridges computational prediction and experimental validation, extending the original in situ framework toward intelligent, feedback-driven bioprocess optimization.







Comments